My research focuses on systems biology approaches to understand quantitatively a minimal bacterium, Mycoplasma pneumoniae. I develop modeling approaches to integrate a whole range of omics data, from genomics to proteomics. The aim of this work is twofold: first, to gain a deeper understanding of the functioning of the cell as a whole, putting the different layers of regulation (transcriptional, translational and post-translational) in the global context of the cell physiology. Second, to serve as a tool in assisting the rational design and development of genetically engineered Mycoplasma cell chassis for synthetic biology applications.
Publications
, 2021. Engineering a genome-reduced bacterium to eliminate Staphylococcus aureus biofilms in vivo. Mol Syst Biol 17(10):e10145
, 2021. Widespread ribosome stalling in a genome-reduced bacterium and the need for translational quality control. iScience 24(9):102985
, 2020. Protein quality control and regulated proteolysis in the genome-reduced organism Mycoplasma pneumoniae. Mol Syst Biol 16(12):e9530
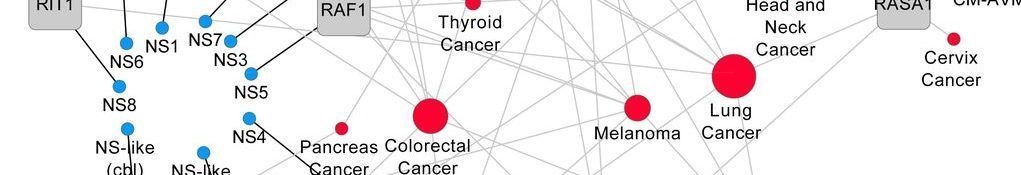
, 2020. Mutation bias within oncogene families is related to proliferation-specific codon usage. Proc Natl Acad Sci U S A 117(48):30848-30856
, 2020. Impact of C-terminal amino acid composition on protein expression in bacteria. Mol Syst Biol 16(5):e9208
, 2016. The cellular Ising model: a framework for phase transitions in multicellular environments. J R Soc Interface 13.
, 2013. Stochastic stabilization of phenotypic States: the genetic bistable switch as a case study. PLoS One 8(9):e73487
, 2013. Dynamics of the quorum sensing switch: stochastic and non-stationary effects. BMC Syst Biol 7:6
, 2011. Noise regulation by quorum sensing in low mRNA copy number systems. BMC Syst Biol 5:11
Publication list retrieved from NCBI using ImpactPubs
.