In our group we are aiming at a quantitative understanding of biological systems to an extent that one is able to predict systemic features and with the hope to rational design and modify their behaviour. This applies to any system comprising biological components that is more than the mere sum of its components, or, in other words, the addition of the individual components results in systemic properties that could not be predicted by considering the components individually. By achieving this objective we are aiming at new global understanding and treatment of human diseases in which the target will not be a single molecule but a network. For this purpose in our group we develop on one hand new software and theoretical approximations to understand complex systems and on the other we do experiments to validate our predictions.
Summary
Latest News
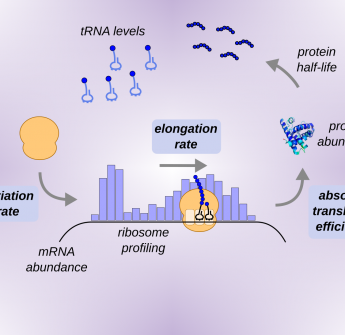
Comprehensive quantitative modeling of translation efficiency in a genome-reduced bacterium
Abstract Translation efficiency has been mainly studied by ribosome profiling, which only provides an incomplete picture of translation kinetics. Here, we integrated the absolute quantifications...
Read More
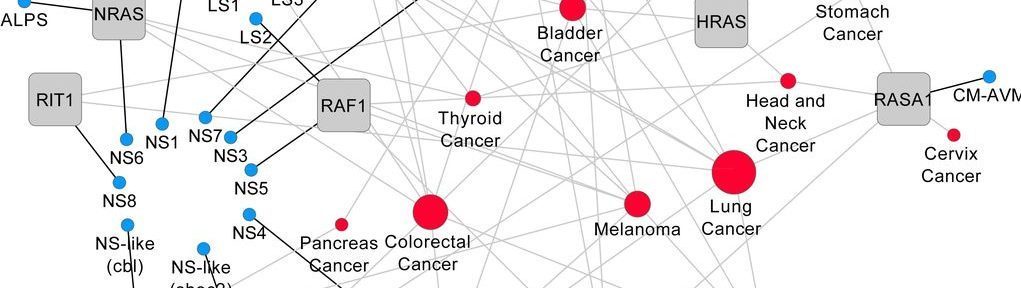
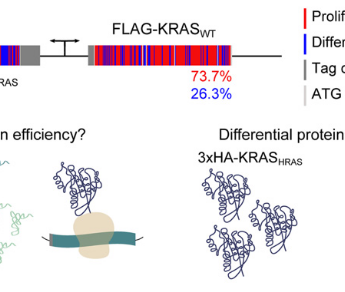
Mutation bias within oncogene families is related to proliferation-specific codon usage
Abstract: "It is well known that in cancer gene families some members are more frequently mutated in tumor samples than their family counterparts. A paradigmatic...
Read More
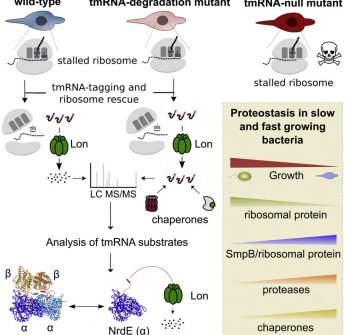
Widespread ribosome stalling in a genome-reduced bacterium and the need for translational quality control
Abstract: "Trans-translation is a ubiquitous bacterial mechanism of ribosome rescue mediated by a transfer-messenger RNA (tmRNA) that adds a degradation tag to the truncated nascent...
Read More
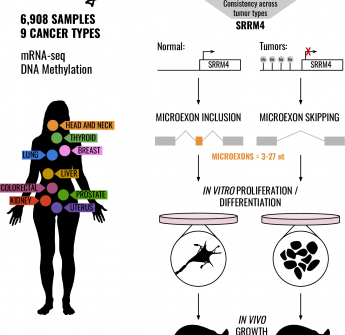
Silencing of SRRM4 suppresses microexon inclusion and promotes tumor growth across cancers
Abstract: "Viruses need to hijack the translational machinery of the host cell for a productive infection to happen. However, given the dynamic landscape of tRNA...
Read More