Abstract:
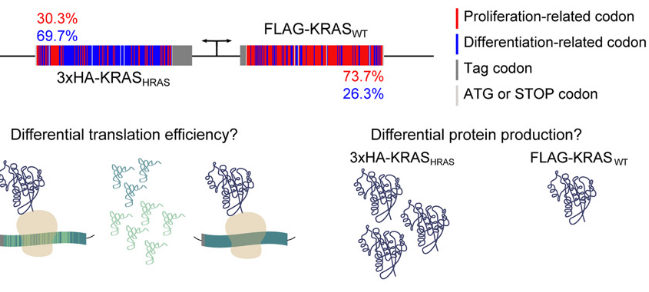
“It is well known that in cancer gene families some members are more frequently mutated in tumor samples than their family counterparts. A paradigmatic case of this phenomenon is KRAS from the RAS family. Different explanations have been proposed ranging from differential interaction with other proteins to preferential expression or localization. Interestingly, it has been described that despite the high amino acid identity between RAS family members, KRAS employs an intriguing differential codon usage. Here, we found that this phenomenon is not exclusive to the RAS family. Indeed, in the RAS family and other oncogene families with two or three members, the most prevalently mutated gene in tumor samples employs a differential codon usage that is characteristic of genes involved in proliferation. Prompted by these observations, we chose the RAS family to experimentally demonstrate that the translation efficiency of oncogenes that are preferentially mutated in tumor samples is increased in proliferative cells compared to quiescent cells. These results were further validated by assessing the translation efficiency of KRAS in cell lines that differ in their tRNA expression profile. These differences are related to the cell division rate of the studied cells and thus suggest an important role in context-specific oncogene expression regulation. Altogether, our study demonstrates that dynamic translation programs contribute to shaping the expression profiles of oncogenes. Therefore, we propose this codon bias as a regulatory layer to control cell context-specific expression and explain the differential prevalence of mutations in certain members of oncogene families.”
Read the full text article.